IntiQuan Webinar Series
Efficient Support of Model Informed Drug Development (MIDD) in R
IntiQuan is holding a series of free webinars based on IntiQuan’s R-based open source modeling & simulation platform
IQR Tools. The topics range from model representation over population modeling, both in the context of PK(PD) and QSP, to advanced clinical trial simulation, considering drop-out and compliance models.
- All material provided (presentation, scripts, models)
- Requirements
- Registration
- Goals / content of the different modules
One of the many advantages of using IQR Tools lies in the fact that a model only needs to be coded once, and this is done in an intuitive format that relies on simple math. Behind the scenes conversion is done in an automatic manner – independent if it is to C-code (for fast simulation) or to NONMEM or MONOLIX (for NLME parameter estimation). In addition, the model syntax is application agnostic, meaning that IQR Tools can support both Pharmacometric and QSP modeling in an identical and equally intuitive manner.
Due to the breadth of topics to cover on the topic of MIDD, we decided to modularize the webinars. This allows for short duration single webinars that allow sequential participation.
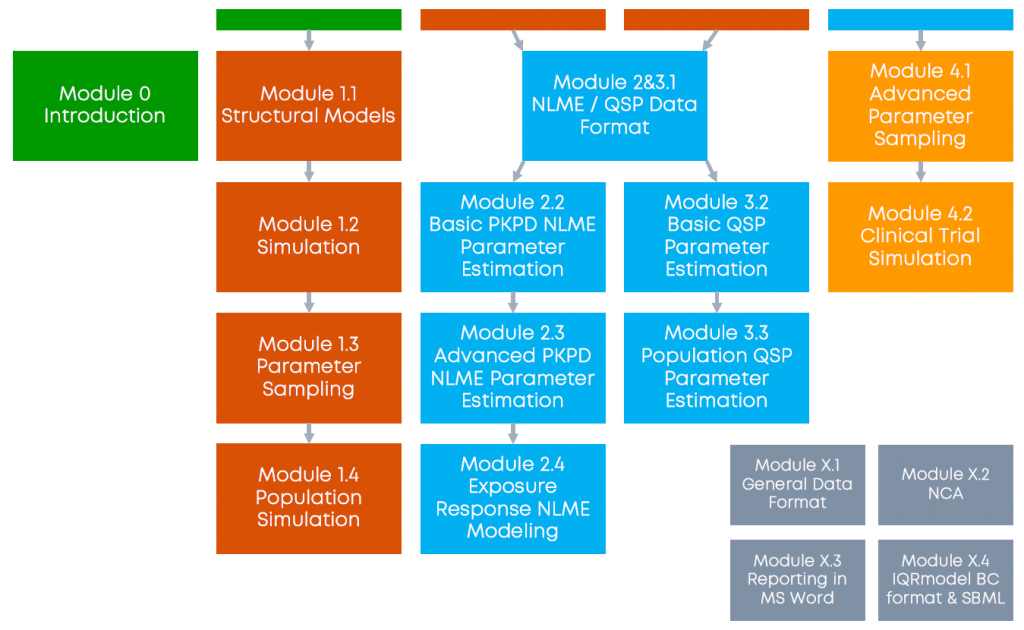
Each webinar module is self contained but assumes (at least some basic) knowledge of the previous modules. QSP and PKPD webinar modules are independent of each other.
Material
Material for each module is provided to participants as a download, including the webinar presentation (PDF) and all examples and scripts that allow to rerun.
Requirements
Not many. You can just watch, listen and get information about state-of-the-art modeling and simulation approaches.
If you would like to run the examples yourself you will need to have an installation of IQR Tools on your computer. Installation is very easy and explained here.
- For Modules 1.x and 4.x you only need R and IQR Tools.
- For Modules 2.x and 3.x you will need in addition an installation of MONOLIX, NONMEM, or NLMIXR on your computer, as in these modules IQR Tools will be used to automatically generate NLME projects, run them, and postprocess results in report ready quality. The setup of tools can be cumbersome (ask your sysadmin about it :-)). And in addition CRAN typically totally messes up reproducibility. For this reason, we suggest to consider the installation of IQdesktop, a freely available virtual computer system that already includes all needed software package for efficient support of MIDD. Just bring you own MONOLIX or NONMEM license (or use NLMIXR). Installation of IQdesktop is easy and typically done within minutes.
Registration
Registration for each webinar module is free of charge but places are limited. You can register from the following webpage: https://training.intiquan.com
Goals of Module 1
| Module | Goals | Time | Link to registration page |
|---|---|---|---|
| M1.1 – Models |
|
90 min | Registration |
| M1.2 – Simulation |
|
90 min | Registration |
| M1.3 – Parameter Sampling |
|
75 min | Registration |
| M1.4 – Population Simulation |
Note: populations simulations go a long way towards clinical trial simulations –this Module 1.4 takes up basic but important concepts. More details on Clinical Trial Simulations are given in Module 4.2”Clinical trial simulation” |
75 min | Registration |
Goals of Module 2
| Module | Goals | Time | Link to registration page |
|---|---|---|---|
| M2.1 – Data Format |
This module is suggested to have seen before embarking on the following modules that will focus on NLME modeling in NONMEM, MONOLIX, and NLMIXR. It will give you the ability to define own datasets in this format. |
60 min | Registration |
| M2.2 – Basic PK/PD (NLME) Parameter estimation |
|
90 min | Registration |
| M2.3 – Advanced PK/PD (NLME) Parameter Estimation |
|
90 min | Registration |
| M2.4 – Exposure Response Modeling |
|
90 min | Registration |
Goals of Module 3
Note that currently not all modules are listed below – this will be updated as the webinar series progresses in 2021.
Goals of Module 4
Note that currently not all modules are listed below – this will be updated as the webinar series progresses in 2021.
Goals of Module X
Note that currently not all modules are listed below – this will be updated as the webinar series progresses in 2021.

